fig = plt.figure(figsize=(22, 14))
s = DISPLAY_STEP
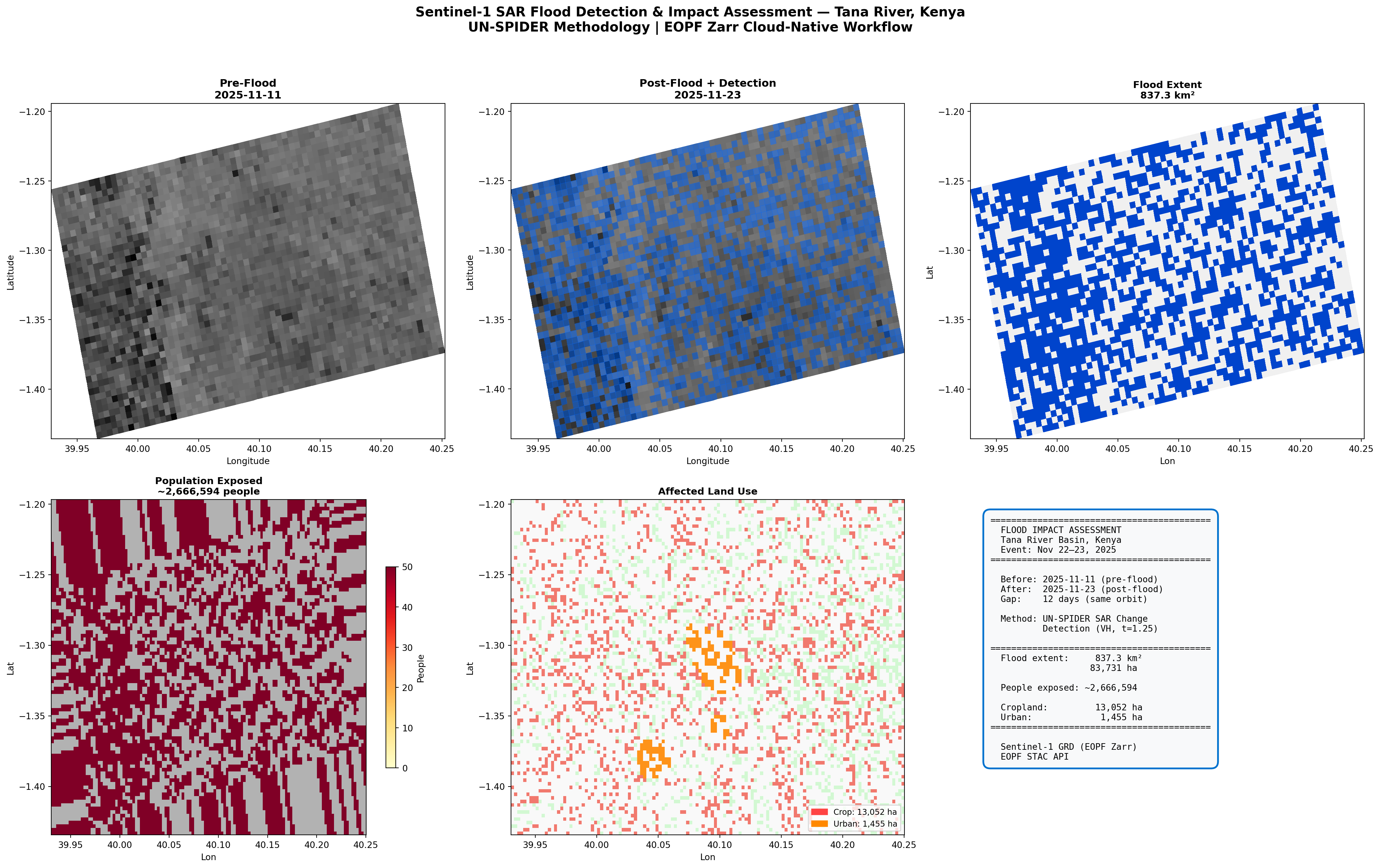
# Top row: SAR imagery and flood detection
ax1 = fig.add_subplot(2, 3, 1)
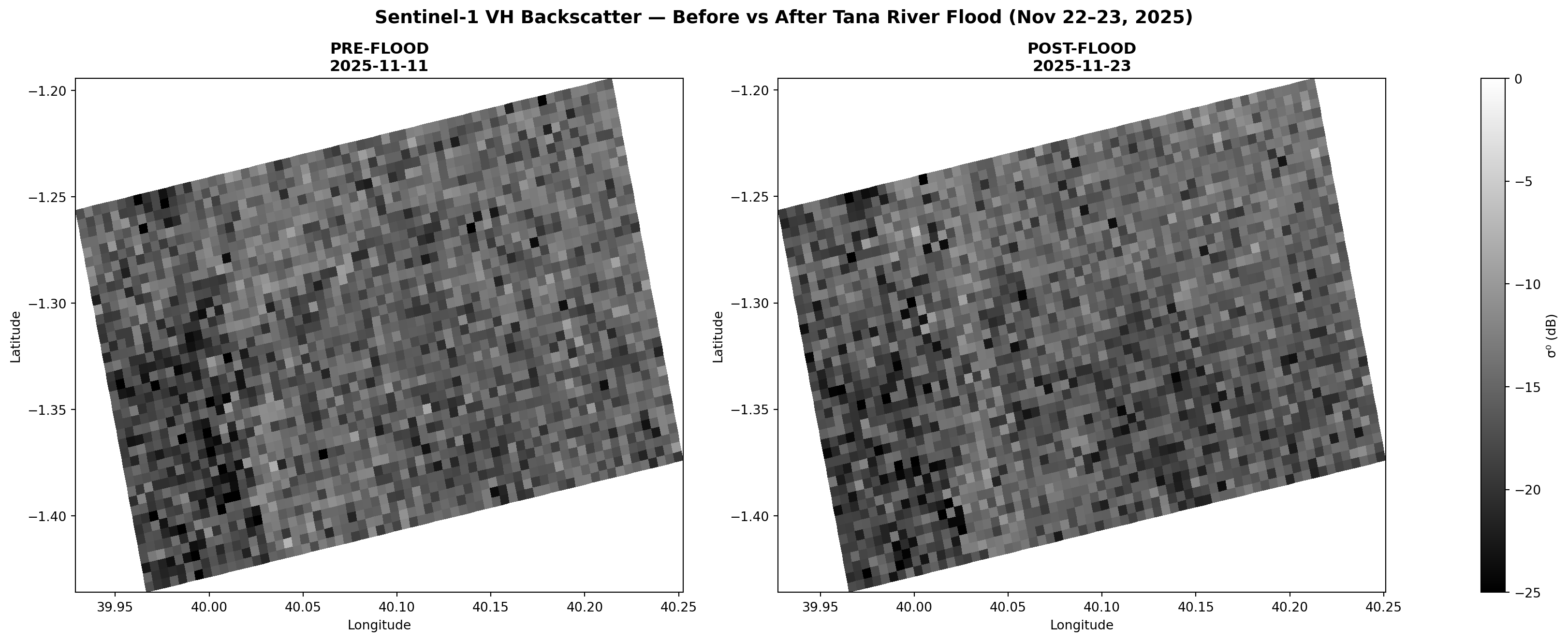
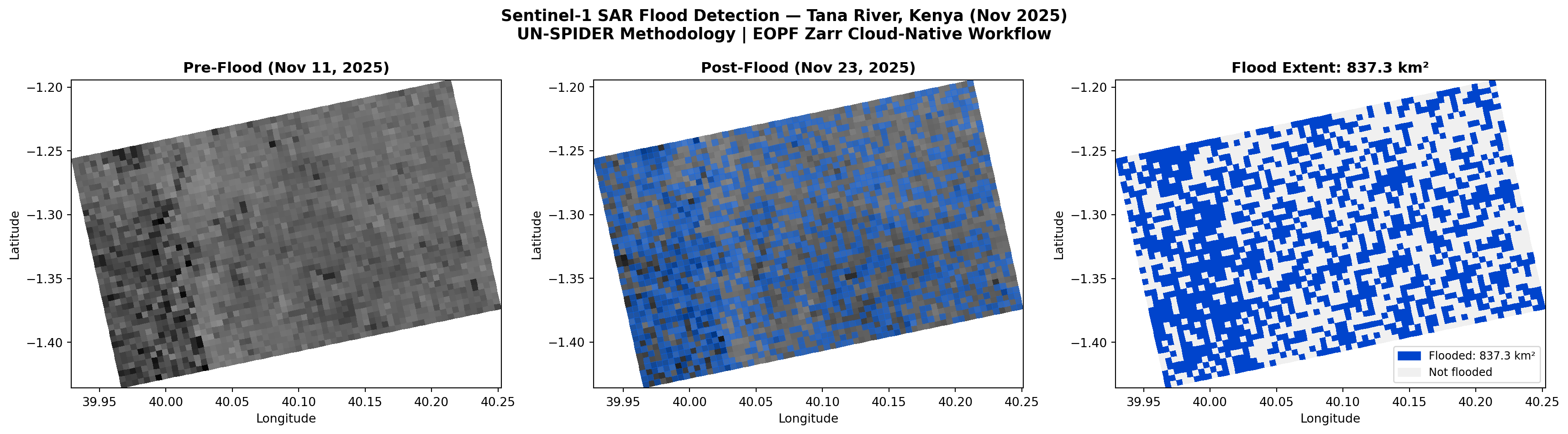
plot_sar(ax1, before_lon, before_lat, before_filt_db, s,
f'Pre-Flood\n{before_item.datetime.strftime("%Y-%m-%d")}',
valid_rows=bvr, valid_cols=bvc)
ax2 = fig.add_subplot(2, 3, 2)
_, alon_c, alat_c = plot_sar(ax2, after_lon, after_lat, after_filt_db, s,
f'Post-Flood + Detection\n{after_item.datetime.strftime("%Y-%m-%d")}',
valid_rows=avr, valid_cols=avc)
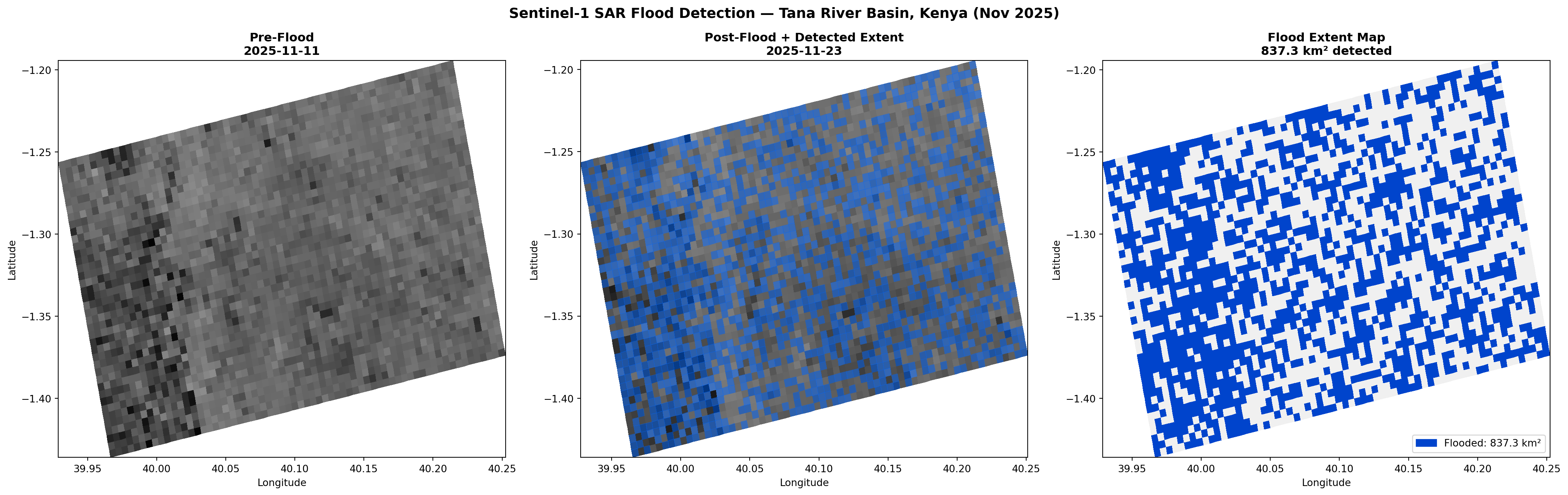
fl_sub = subsample_to_coords(flood_refined, s, after_lat, avr, avc)
fl_ma = np.ma.masked_where(~fl_sub, fl_sub.astype(float))
ax2.pcolormesh(alon_c, alat_c, fl_ma, cmap=ListedColormap(['#0066FF']), alpha=0.5)
ax3 = fig.add_subplot(2, 3, 3)
fl_sub3 = subsample_to_coords(flood_refined, s, before_lat, bvr, bvc)
_, blon_c, blat_c = plot_sar(ax3, before_lon, before_lat, before_filt_db, s, '',
valid_rows=bvr, valid_cols=bvc)
ax3.clear()
ax3.pcolormesh(blon_c, blat_c, fl_sub3.astype(int),
cmap=ListedColormap(['#F0F0F0','#0044CC']), vmin=0, vmax=1)
ax3.set_title(f'Flood Extent\n{flood_km2:,.1f} km²', fontsize=11, fontweight='bold')
ax3.set_xlabel('Lon')
ax3.set_ylabel('Lat')
# Bottom row: Impact assessment
ax4 = fig.add_subplot(2, 3, 4)
ax4.pcolormesh(rlon, rlat, pop, cmap='Greys', vmin=0, vmax=100, alpha=0.3)
pe = np.ma.masked_where(~flood_reg, pop_exposed)
im4 = ax4.pcolormesh(rlon, rlat, pe, cmap='YlOrRd', vmin=0, vmax=50)
fig.colorbar(im4, ax=ax4, label='People', shrink=0.6)
ax4.set_title(f'Population Exposed\n~{total_exposed:,} people', fontsize=11, fontweight='bold')
ax4.set_xlabel('Lon')
ax4.set_ylabel('Lat')
ax5 = fig.add_subplot(2, 3, 5)
ax5.pcolormesh(rlon, rlat, ((lc==12)|(lc==14)).astype(float),
cmap=ListedColormap(['#F0F0F0','#90EE90']), vmin=0, vmax=1, alpha=0.4)
ac = np.ma.masked_where(~crop_affected, crop_affected.astype(float))
ax5.pcolormesh(rlon, rlat, ac, cmap=ListedColormap(['#FF4444']), alpha=0.7)
au = np.ma.masked_where(~urban_affected, urban_affected.astype(float))
ax5.pcolormesh(rlon, rlat, au, cmap=ListedColormap(['#FF8800']), alpha=0.9)
ax5.legend(handles=[
mpatches.Patch(color='#FF4444', label=f'Crop: {crop_ha:,} ha'),
mpatches.Patch(color='#FF8800', label=f'Urban: {urban_ha:,} ha')
], loc='lower right', fontsize=9)
ax5.set_title('Affected Land Use', fontsize=11, fontweight='bold')
ax5.set_xlabel('Lon')
ax5.set_ylabel('Lat')
# Summary panel
ax6 = fig.add_subplot(2, 3, 6)
ax6.axis('off')
summary = (
f"{'='*42}\n"
f" FLOOD IMPACT ASSESSMENT\n"
f" Tana River Basin, Kenya\n"
f" Event: Nov 22–23, 2025\n"
f"{'='*42}\n\n"
f" Before: {before_item.datetime.strftime('%Y-%m-%d')} (pre-flood)\n"
f" After: {after_item.datetime.strftime('%Y-%m-%d')} (post-flood)\n"
f" Gap: 12 days (same orbit)\n\n"
f" Method: UN-SPIDER SAR Change\n"
f" Detection (VH, t={effective_threshold})\n\n"
f"{'='*42}\n"
f" Flood extent: {flood_km2:>8,.1f} km²\n"
f" {flood_ha:>8,.0f} ha\n\n"
f" People exposed: ~{total_exposed:>7,}\n\n"
f" Cropland: {crop_ha:>8,} ha\n"
f" Urban: {urban_ha:>8,} ha\n"
f"{'='*42}\n\n"
f" Sentinel-1 GRD (EOPF Zarr)\n"
f" EOPF STAC API"
)
ax6.text(0.05, 0.95, summary, transform=ax6.transAxes, fontsize=10,
fontfamily='monospace', va='top',
bbox=dict(boxstyle='round,pad=0.8', fc='#f8f9fa', ec='#0072ce', lw=2))
fig.suptitle('Sentinel-1 SAR Flood Detection & Impact Assessment — Tana River, Kenya\n'
'UN-SPIDER Methodology | EOPF Zarr Cloud-Native Workflow',
fontsize=15, fontweight='bold', y=0.98)
plt.tight_layout(rect=[0,0,1,0.95])
plt.show()