zarr_url = "https://objects.eodc.eu/e05ab01a9d56408d82ac32d69a5aae2a:sample-data/tutorial_data/cpm_v253/S2B_MSIL1C_20250113T103309_N0511_R108_T32TLQ_20250113T122458.zarr"Access EOPF Sentinel Zarr using Rarr
By: @joheisig | Institut für Geoinformatik
Introduction
This notebook demonstrates how to retrieve remotely stored Zarr data using the Rarr package in R. We will explore how to read and visualize Zarr data (zarrays) and their metadata. For subsequent analysis and visualization we turn the zarray into stars objects which are more suitable for spatial data.
This notebook has a sibling, which demonstrates how to access the same data using the GDAL Zarr driver. You can find it here.
What we will learn
- ✏️ How to edit URLs of Zarr stores to make them readable for GDAL
- 🔎 Which read-functions and arguments to use in
starsandterra - 🚧 Current limitations of these packages
Prerequisites
To follow this notebook, you will need to load a STAC item in Zarr format from the EOPF Zarr STAC Catalog. If you are new to STAC inside the R environment, we suggest to follow the Access the EOPF Zarr STAC API with R tutorial. The item we are using for this example mirrors the one used in the Sentinel-2 L1C MSI Zarr Product Exploration notebook which is hosted on EODC’s object storage and is accessible under the URL below.
Import packages
library(Rarr) # reading; BiocManager::install("Rarr")
library(stars) # spatial objects
library(dplyr) # table operations
library(jsonlite) # parsing metadata
library(mapview) # interactive mapsMetadata exploration with Rarr
In this notebook we show the current capabilities and limitations of Rarr in terms of reading Zarr array(s) and their metadata. Successful attempts are marked with a ✅, failed attempts with a ⛔.
Using the Rarr::zarr_overview function we can get meta data for all assets in the remote Zarr store as a table. Here list all arrays and their data type, compression, dimensions, etc.
zarr_content = zarr_overview(zarr_url, as_data_frame = T) |>
mutate(var = sub(zarr_url, "", path)) |>
select(var, everything(), -path)
filter(zarr_content, grepl("reflectance", var)) var data_type endianness compressor
1 /measurements/reflectance/r10m/b02 uint16 little blosc
2 /measurements/reflectance/r10m/b03 uint16 little blosc
3 /measurements/reflectance/r10m/b04 uint16 little blosc
4 /measurements/reflectance/r10m/b08 uint16 little blosc
5 /measurements/reflectance/r10m/x int64 little blosc
6 /measurements/reflectance/r10m/y int64 little blosc
7 /measurements/reflectance/r20m/b05 uint16 little blosc
8 /measurements/reflectance/r20m/b06 uint16 little blosc
9 /measurements/reflectance/r20m/b07 uint16 little blosc
10 /measurements/reflectance/r20m/b11 uint16 little blosc
11 /measurements/reflectance/r20m/b12 uint16 little blosc
12 /measurements/reflectance/r20m/b8a uint16 little blosc
13 /measurements/reflectance/r20m/x int64 little blosc
14 /measurements/reflectance/r20m/y int64 little blosc
15 /measurements/reflectance/r60m/b01 uint16 little blosc
16 /measurements/reflectance/r60m/b09 uint16 little blosc
17 /measurements/reflectance/r60m/b10 uint16 little blosc
18 /measurements/reflectance/r60m/x int64 little blosc
19 /measurements/reflectance/r60m/y int64 little blosc
dim chunk_dim nchunks
1 10980, 10980 1830, 1830 6, 6
2 10980, 10980 1830, 1830 6, 6
3 10980, 10980 1830, 1830 6, 6
4 10980, 10980 1830, 1830 6, 6
5 10980 10980 1
6 10980 10980 1
7 5490, 5490 915, 915 6, 6
8 5490, 5490 915, 915 6, 6
9 5490, 5490 915, 915 6, 6
10 5490, 5490 915, 915 6, 6
11 5490, 5490 915, 915 6, 6
12 5490, 5490 915, 915 6, 6
13 5490 5490 1
14 5490 5490 1
15 1830, 1830 305, 305 6, 6
16 1830, 1830 305, 305 6, 6
17 1830, 1830 305, 305 6, 6
18 1830 1830 1
19 1830 1830 1We can see that all Sentinel-2 bands are present, but split into directories (or groups) based on spatial resolution (10, 20, 60 meters), which have different dimensions (number and size of pixels).
var data_type endianness
1 /conditions/geometry/angle unicode224 little
2 /conditions/geometry/band unicode96 little
3 /conditions/geometry/detector int64 little
4 /conditions/geometry/mean_sun_angles float64 little
5 /conditions/geometry/mean_viewing_incidence_angles float64 little
6 /conditions/geometry/sun_angles float64 little
7 /conditions/geometry/viewing_incidence_angles float64 little
8 /conditions/geometry/x int64 little
9 /conditions/geometry/y int64 little
10 /conditions/mask/detector_footprint/r10m/b02 uint8 <NA>
11 /conditions/mask/detector_footprint/r10m/b03 uint8 <NA>
12 /conditions/mask/detector_footprint/r10m/b04 uint8 <NA>
13 /conditions/mask/detector_footprint/r10m/b08 uint8 <NA>
14 /conditions/mask/detector_footprint/r10m/x int64 little
15 /conditions/mask/detector_footprint/r10m/y int64 little
16 /conditions/mask/detector_footprint/r20m/b05 uint8 <NA>
17 /conditions/mask/detector_footprint/r20m/b06 uint8 <NA>
18 /conditions/mask/detector_footprint/r20m/b07 uint8 <NA>
19 /conditions/mask/detector_footprint/r20m/b11 uint8 <NA>
20 /conditions/mask/detector_footprint/r20m/b12 uint8 <NA>
21 /conditions/mask/detector_footprint/r20m/b8a uint8 <NA>
22 /conditions/mask/detector_footprint/r20m/x int64 little
23 /conditions/mask/detector_footprint/r20m/y int64 little
24 /conditions/mask/detector_footprint/r60m/b01 uint8 <NA>
25 /conditions/mask/detector_footprint/r60m/b09 uint8 <NA>
26 /conditions/mask/detector_footprint/r60m/b10 uint8 <NA>
27 /conditions/mask/detector_footprint/r60m/x int64 little
28 /conditions/mask/detector_footprint/r60m/y int64 little
29 /conditions/mask/l1c_classification/r60m/b00 uint8 <NA>
30 /conditions/mask/l1c_classification/r60m/x int64 little
31 /conditions/mask/l1c_classification/r60m/y int64 little
32 /conditions/meteorology/cams/aod1240 float32 little
33 /conditions/meteorology/cams/aod469 float32 little
34 /conditions/meteorology/cams/aod550 float32 little
35 /conditions/meteorology/cams/aod670 float32 little
36 /conditions/meteorology/cams/aod865 float32 little
37 /conditions/meteorology/cams/bcaod550 float32 little
38 /conditions/meteorology/cams/duaod550 float32 little
39 /conditions/meteorology/cams/isobaricInhPa float64 little
40 /conditions/meteorology/cams/latitude float64 little
41 /conditions/meteorology/cams/longitude float64 little
42 /conditions/meteorology/cams/number int64 little
43 /conditions/meteorology/cams/omaod550 float32 little
44 /conditions/meteorology/cams/ssaod550 float32 little
45 /conditions/meteorology/cams/step int64 little
46 /conditions/meteorology/cams/suaod550 float32 little
47 /conditions/meteorology/cams/surface float64 little
48 /conditions/meteorology/cams/time int64 little
49 /conditions/meteorology/cams/valid_time int64 little
50 /conditions/meteorology/cams/z float32 little
51 /conditions/meteorology/ecmwf/isobaricInhPa float64 little
52 /conditions/meteorology/ecmwf/latitude float64 little
53 /conditions/meteorology/ecmwf/longitude float64 little
54 /conditions/meteorology/ecmwf/msl float32 little
55 /conditions/meteorology/ecmwf/number int64 little
56 /conditions/meteorology/ecmwf/r float32 little
57 /conditions/meteorology/ecmwf/step int64 little
58 /conditions/meteorology/ecmwf/surface float64 little
59 /conditions/meteorology/ecmwf/tco3 float32 little
60 /conditions/meteorology/ecmwf/tcwv float32 little
61 /conditions/meteorology/ecmwf/time int64 little
62 /conditions/meteorology/ecmwf/u10 float32 little
63 /conditions/meteorology/ecmwf/v10 float32 little
64 /conditions/meteorology/ecmwf/valid_time int64 little
65 /measurements/reflectance/r10m/b02 uint16 little
66 /measurements/reflectance/r10m/b03 uint16 little
67 /measurements/reflectance/r10m/b04 uint16 little
68 /measurements/reflectance/r10m/b08 uint16 little
69 /measurements/reflectance/r10m/x int64 little
70 /measurements/reflectance/r10m/y int64 little
71 /measurements/reflectance/r20m/b05 uint16 little
72 /measurements/reflectance/r20m/b06 uint16 little
73 /measurements/reflectance/r20m/b07 uint16 little
74 /measurements/reflectance/r20m/b11 uint16 little
75 /measurements/reflectance/r20m/b12 uint16 little
76 /measurements/reflectance/r20m/b8a uint16 little
77 /measurements/reflectance/r20m/x int64 little
78 /measurements/reflectance/r20m/y int64 little
79 /measurements/reflectance/r60m/b01 uint16 little
80 /measurements/reflectance/r60m/b09 uint16 little
81 /measurements/reflectance/r60m/b10 uint16 little
82 /measurements/reflectance/r60m/x int64 little
83 /measurements/reflectance/r60m/y int64 little
84 /quality/l1c_quicklook/r10m/band int64 little
85 /quality/l1c_quicklook/r10m/tci uint8 <NA>
86 /quality/l1c_quicklook/r10m/x int64 little
87 /quality/l1c_quicklook/r10m/y int64 little
88 /quality/mask/r10m/b02 uint8 <NA>
89 /quality/mask/r10m/b03 uint8 <NA>
90 /quality/mask/r10m/b04 uint8 <NA>
91 /quality/mask/r10m/b08 uint8 <NA>
92 /quality/mask/r10m/x int64 little
93 /quality/mask/r10m/y int64 little
94 /quality/mask/r20m/b05 uint8 <NA>
95 /quality/mask/r20m/b06 uint8 <NA>
96 /quality/mask/r20m/b07 uint8 <NA>
97 /quality/mask/r20m/b11 uint8 <NA>
98 /quality/mask/r20m/b12 uint8 <NA>
99 /quality/mask/r20m/b8a uint8 <NA>
100 /quality/mask/r20m/x int64 little
101 /quality/mask/r20m/y int64 little
102 /quality/mask/r60m/b01 uint8 <NA>
103 /quality/mask/r60m/b09 uint8 <NA>
104 /quality/mask/r60m/b10 uint8 <NA>
105 /quality/mask/r60m/x int64 little
106 /quality/mask/r60m/y int64 little
compressor dim chunk_dim nchunks
1 blosc 2 2 1
2 blosc 13 13 1
3 blosc 7 7 1
4 blosc 2 2 1
5 blosc 13, 2 13, 2 1, 1
6 blosc 2, 23, 23 2, 23, 23 1, 1, 1
7 blosc 13, 7, 2.... 7, 4, 2,.... 2, 2, 1,....
8 blosc 23 23 1
9 blosc 23 23 1
10 blosc 10980, 10980 1830, 1830 6, 6
11 blosc 10980, 10980 1830, 1830 6, 6
12 blosc 10980, 10980 1830, 1830 6, 6
13 blosc 10980, 10980 1830, 1830 6, 6
14 blosc 10980 10980 1
15 blosc 10980 10980 1
16 blosc 5490, 5490 915, 915 6, 6
17 blosc 5490, 5490 915, 915 6, 6
18 blosc 5490, 5490 915, 915 6, 6
19 blosc 5490, 5490 915, 915 6, 6
20 blosc 5490, 5490 915, 915 6, 6
21 blosc 5490, 5490 915, 915 6, 6
22 blosc 5490 5490 1
23 blosc 5490 5490 1
24 blosc 1830, 1830 305, 305 6, 6
25 blosc 1830, 1830 305, 305 6, 6
26 blosc 1830, 1830 305, 305 6, 6
27 blosc 1830 1830 1
28 blosc 1830 1830 1
29 blosc 1830, 1830 305, 305 6, 6
30 blosc 1830 1830 1
31 blosc 1830 1830 1
32 blosc 9, 9 9, 9 1, 1
33 blosc 9, 9 9, 9 1, 1
34 blosc 9, 9 9, 9 1, 1
35 blosc 9, 9 9, 9 1, 1
36 blosc 9, 9 9, 9 1, 1
37 blosc 9, 9 9, 9 1, 1
38 blosc 9, 9 9, 9 1, 1
39 <NA>
40 blosc 9 9 1
41 blosc 9 9 1
42 <NA>
43 blosc 9, 9 9, 9 1, 1
44 blosc 9, 9 9, 9 1, 1
45 <NA>
46 blosc 9, 9 9, 9 1, 1
47 <NA>
48 <NA>
49 <NA>
50 blosc 9, 9 9, 9 1, 1
51 <NA>
52 blosc 9 9 1
53 blosc 9 9 1
54 blosc 9, 9 9, 9 1, 1
55 <NA>
56 blosc 9, 9 9, 9 1, 1
57 <NA>
58 <NA>
59 blosc 9, 9 9, 9 1, 1
60 blosc 9, 9 9, 9 1, 1
61 <NA>
62 blosc 9, 9 9, 9 1, 1
63 blosc 9, 9 9, 9 1, 1
64 <NA>
65 blosc 10980, 10980 1830, 1830 6, 6
66 blosc 10980, 10980 1830, 1830 6, 6
67 blosc 10980, 10980 1830, 1830 6, 6
68 blosc 10980, 10980 1830, 1830 6, 6
69 blosc 10980 10980 1
70 blosc 10980 10980 1
71 blosc 5490, 5490 915, 915 6, 6
72 blosc 5490, 5490 915, 915 6, 6
73 blosc 5490, 5490 915, 915 6, 6
74 blosc 5490, 5490 915, 915 6, 6
75 blosc 5490, 5490 915, 915 6, 6
76 blosc 5490, 5490 915, 915 6, 6
77 blosc 5490 5490 1
78 blosc 5490 5490 1
79 blosc 1830, 1830 305, 305 6, 6
80 blosc 1830, 1830 305, 305 6, 6
81 blosc 1830, 1830 305, 305 6, 6
82 blosc 1830 1830 1
83 blosc 1830 1830 1
84 blosc 3 3 1
85 blosc 3, 10980.... 1, 1830,.... 3, 6, 6
86 blosc 10980 10980 1
87 blosc 10980 10980 1
88 blosc 10980, 10980 1830, 1830 6, 6
89 blosc 10980, 10980 1830, 1830 6, 6
90 blosc 10980, 10980 1830, 1830 6, 6
91 blosc 10980, 10980 1830, 1830 6, 6
92 blosc 10980 10980 1
93 blosc 10980 10980 1
94 blosc 5490, 5490 915, 915 6, 6
95 blosc 5490, 5490 915, 915 6, 6
96 blosc 5490, 5490 915, 915 6, 6
97 blosc 5490, 5490 915, 915 6, 6
98 blosc 5490, 5490 915, 915 6, 6
99 blosc 5490, 5490 915, 915 6, 6
100 blosc 5490 5490 1
101 blosc 5490 5490 1
102 blosc 1830, 1830 305, 305 6, 6
103 blosc 1830, 1830 305, 305 6, 6
104 blosc 1830, 1830 305, 305 6, 6
105 blosc 1830 1830 1
106 blosc 1830 1830 1Now that we have explored the content, let’s get some related metadata. Rarr provides the read_zarr_attributes function for this purpose, which looks for the .zattrs file inside the Zarr store.
⛔ Unfortunately we were not successful when applying it to our remote store.
read_zarr_attributes(zarr_url)Error:
! No file that could contain attributes (either `.zattrs` for v2 or `zarr.json` for v3) was found in the path.read_zarr_attributes(file.path(zarr_url, ".zattrs"))Error:
! No file that could contain attributes (either `.zattrs` for v2 or `zarr.json` for v3) was found in the path.✅ As a simple workaround we can read the JSON directly:
zattrs = read_json(file.path(zarr_url, ".zattrs"))
names(zattrs)[1] "other_metadata" "other_metadata4" "stac_discovery" var data_type endianness
1 /conditions/geometry/angle unicode224 little
2 /conditions/geometry/band unicode96 little
3 /conditions/geometry/detector int64 little
4 /conditions/geometry/mean_sun_angles float64 little
5 /conditions/geometry/mean_viewing_incidence_angles float64 little
6 /conditions/geometry/sun_angles float64 little
7 /conditions/geometry/viewing_incidence_angles float64 little
8 /conditions/geometry/x int64 little
9 /conditions/geometry/y int64 little
10 /conditions/mask/detector_footprint/r10m/b02 uint8 <NA>
11 /conditions/mask/detector_footprint/r10m/b03 uint8 <NA>
12 /conditions/mask/detector_footprint/r10m/b04 uint8 <NA>
13 /conditions/mask/detector_footprint/r10m/b08 uint8 <NA>
14 /conditions/mask/detector_footprint/r10m/x int64 little
15 /conditions/mask/detector_footprint/r10m/y int64 little
16 /conditions/mask/detector_footprint/r20m/b05 uint8 <NA>
17 /conditions/mask/detector_footprint/r20m/b06 uint8 <NA>
18 /conditions/mask/detector_footprint/r20m/b07 uint8 <NA>
19 /conditions/mask/detector_footprint/r20m/b11 uint8 <NA>
20 /conditions/mask/detector_footprint/r20m/b12 uint8 <NA>
21 /conditions/mask/detector_footprint/r20m/b8a uint8 <NA>
22 /conditions/mask/detector_footprint/r20m/x int64 little
23 /conditions/mask/detector_footprint/r20m/y int64 little
24 /conditions/mask/detector_footprint/r60m/b01 uint8 <NA>
25 /conditions/mask/detector_footprint/r60m/b09 uint8 <NA>
26 /conditions/mask/detector_footprint/r60m/b10 uint8 <NA>
27 /conditions/mask/detector_footprint/r60m/x int64 little
28 /conditions/mask/detector_footprint/r60m/y int64 little
29 /conditions/mask/l1c_classification/r60m/b00 uint8 <NA>
30 /conditions/mask/l1c_classification/r60m/x int64 little
31 /conditions/mask/l1c_classification/r60m/y int64 little
32 /conditions/meteorology/cams/aod1240 float32 little
33 /conditions/meteorology/cams/aod469 float32 little
34 /conditions/meteorology/cams/aod550 float32 little
35 /conditions/meteorology/cams/aod670 float32 little
36 /conditions/meteorology/cams/aod865 float32 little
37 /conditions/meteorology/cams/bcaod550 float32 little
38 /conditions/meteorology/cams/duaod550 float32 little
39 /conditions/meteorology/cams/isobaricInhPa float64 little
40 /conditions/meteorology/cams/latitude float64 little
41 /conditions/meteorology/cams/longitude float64 little
42 /conditions/meteorology/cams/number int64 little
43 /conditions/meteorology/cams/omaod550 float32 little
44 /conditions/meteorology/cams/ssaod550 float32 little
45 /conditions/meteorology/cams/step int64 little
46 /conditions/meteorology/cams/suaod550 float32 little
47 /conditions/meteorology/cams/surface float64 little
48 /conditions/meteorology/cams/time int64 little
49 /conditions/meteorology/cams/valid_time int64 little
50 /conditions/meteorology/cams/z float32 little
51 /conditions/meteorology/ecmwf/isobaricInhPa float64 little
52 /conditions/meteorology/ecmwf/latitude float64 little
53 /conditions/meteorology/ecmwf/longitude float64 little
54 /conditions/meteorology/ecmwf/msl float32 little
55 /conditions/meteorology/ecmwf/number int64 little
56 /conditions/meteorology/ecmwf/r float32 little
57 /conditions/meteorology/ecmwf/step int64 little
58 /conditions/meteorology/ecmwf/surface float64 little
59 /conditions/meteorology/ecmwf/tco3 float32 little
60 /conditions/meteorology/ecmwf/tcwv float32 little
61 /conditions/meteorology/ecmwf/time int64 little
62 /conditions/meteorology/ecmwf/u10 float32 little
63 /conditions/meteorology/ecmwf/v10 float32 little
64 /conditions/meteorology/ecmwf/valid_time int64 little
65 /measurements/reflectance/r10m/b02 uint16 little
66 /measurements/reflectance/r10m/b03 uint16 little
67 /measurements/reflectance/r10m/b04 uint16 little
68 /measurements/reflectance/r10m/b08 uint16 little
69 /measurements/reflectance/r10m/x int64 little
70 /measurements/reflectance/r10m/y int64 little
71 /measurements/reflectance/r20m/b05 uint16 little
72 /measurements/reflectance/r20m/b06 uint16 little
73 /measurements/reflectance/r20m/b07 uint16 little
74 /measurements/reflectance/r20m/b11 uint16 little
75 /measurements/reflectance/r20m/b12 uint16 little
76 /measurements/reflectance/r20m/b8a uint16 little
77 /measurements/reflectance/r20m/x int64 little
78 /measurements/reflectance/r20m/y int64 little
79 /measurements/reflectance/r60m/b01 uint16 little
80 /measurements/reflectance/r60m/b09 uint16 little
81 /measurements/reflectance/r60m/b10 uint16 little
82 /measurements/reflectance/r60m/x int64 little
83 /measurements/reflectance/r60m/y int64 little
84 /quality/l1c_quicklook/r10m/band int64 little
85 /quality/l1c_quicklook/r10m/tci uint8 <NA>
86 /quality/l1c_quicklook/r10m/x int64 little
87 /quality/l1c_quicklook/r10m/y int64 little
88 /quality/mask/r10m/b02 uint8 <NA>
89 /quality/mask/r10m/b03 uint8 <NA>
90 /quality/mask/r10m/b04 uint8 <NA>
91 /quality/mask/r10m/b08 uint8 <NA>
92 /quality/mask/r10m/x int64 little
93 /quality/mask/r10m/y int64 little
94 /quality/mask/r20m/b05 uint8 <NA>
95 /quality/mask/r20m/b06 uint8 <NA>
96 /quality/mask/r20m/b07 uint8 <NA>
97 /quality/mask/r20m/b11 uint8 <NA>
98 /quality/mask/r20m/b12 uint8 <NA>
99 /quality/mask/r20m/b8a uint8 <NA>
100 /quality/mask/r20m/x int64 little
101 /quality/mask/r20m/y int64 little
102 /quality/mask/r60m/b01 uint8 <NA>
103 /quality/mask/r60m/b09 uint8 <NA>
104 /quality/mask/r60m/b10 uint8 <NA>
105 /quality/mask/r60m/x int64 little
106 /quality/mask/r60m/y int64 little
compressor dim chunk_dim nchunks
1 blosc 2 2 1
2 blosc 13 13 1
3 blosc 7 7 1
4 blosc 2 2 1
5 blosc 13, 2 13, 2 1, 1
6 blosc 2, 23, 23 2, 23, 23 1, 1, 1
7 blosc 13, 7, 2.... 7, 4, 2,.... 2, 2, 1,....
8 blosc 23 23 1
9 blosc 23 23 1
10 blosc 10980, 10980 1830, 1830 6, 6
11 blosc 10980, 10980 1830, 1830 6, 6
12 blosc 10980, 10980 1830, 1830 6, 6
13 blosc 10980, 10980 1830, 1830 6, 6
14 blosc 10980 10980 1
15 blosc 10980 10980 1
16 blosc 5490, 5490 915, 915 6, 6
17 blosc 5490, 5490 915, 915 6, 6
18 blosc 5490, 5490 915, 915 6, 6
19 blosc 5490, 5490 915, 915 6, 6
20 blosc 5490, 5490 915, 915 6, 6
21 blosc 5490, 5490 915, 915 6, 6
22 blosc 5490 5490 1
23 blosc 5490 5490 1
24 blosc 1830, 1830 305, 305 6, 6
25 blosc 1830, 1830 305, 305 6, 6
26 blosc 1830, 1830 305, 305 6, 6
27 blosc 1830 1830 1
28 blosc 1830 1830 1
29 blosc 1830, 1830 305, 305 6, 6
30 blosc 1830 1830 1
31 blosc 1830 1830 1
32 blosc 9, 9 9, 9 1, 1
33 blosc 9, 9 9, 9 1, 1
34 blosc 9, 9 9, 9 1, 1
35 blosc 9, 9 9, 9 1, 1
36 blosc 9, 9 9, 9 1, 1
37 blosc 9, 9 9, 9 1, 1
38 blosc 9, 9 9, 9 1, 1
39 <NA>
40 blosc 9 9 1
41 blosc 9 9 1
42 <NA>
43 blosc 9, 9 9, 9 1, 1
44 blosc 9, 9 9, 9 1, 1
45 <NA>
46 blosc 9, 9 9, 9 1, 1
47 <NA>
48 <NA>
49 <NA>
50 blosc 9, 9 9, 9 1, 1
51 <NA>
52 blosc 9 9 1
53 blosc 9 9 1
54 blosc 9, 9 9, 9 1, 1
55 <NA>
56 blosc 9, 9 9, 9 1, 1
57 <NA>
58 <NA>
59 blosc 9, 9 9, 9 1, 1
60 blosc 9, 9 9, 9 1, 1
61 <NA>
62 blosc 9, 9 9, 9 1, 1
63 blosc 9, 9 9, 9 1, 1
64 <NA>
65 blosc 10980, 10980 1830, 1830 6, 6
66 blosc 10980, 10980 1830, 1830 6, 6
67 blosc 10980, 10980 1830, 1830 6, 6
68 blosc 10980, 10980 1830, 1830 6, 6
69 blosc 10980 10980 1
70 blosc 10980 10980 1
71 blosc 5490, 5490 915, 915 6, 6
72 blosc 5490, 5490 915, 915 6, 6
73 blosc 5490, 5490 915, 915 6, 6
74 blosc 5490, 5490 915, 915 6, 6
75 blosc 5490, 5490 915, 915 6, 6
76 blosc 5490, 5490 915, 915 6, 6
77 blosc 5490 5490 1
78 blosc 5490 5490 1
79 blosc 1830, 1830 305, 305 6, 6
80 blosc 1830, 1830 305, 305 6, 6
81 blosc 1830, 1830 305, 305 6, 6
82 blosc 1830 1830 1
83 blosc 1830 1830 1
84 blosc 3 3 1
85 blosc 3, 10980.... 1, 1830,.... 3, 6, 6
86 blosc 10980 10980 1
87 blosc 10980 10980 1
88 blosc 10980, 10980 1830, 1830 6, 6
89 blosc 10980, 10980 1830, 1830 6, 6
90 blosc 10980, 10980 1830, 1830 6, 6
91 blosc 10980, 10980 1830, 1830 6, 6
92 blosc 10980 10980 1
93 blosc 10980 10980 1
94 blosc 5490, 5490 915, 915 6, 6
95 blosc 5490, 5490 915, 915 6, 6
96 blosc 5490, 5490 915, 915 6, 6
97 blosc 5490, 5490 915, 915 6, 6
98 blosc 5490, 5490 915, 915 6, 6
99 blosc 5490, 5490 915, 915 6, 6
100 blosc 5490 5490 1
101 blosc 5490 5490 1
102 blosc 1830, 1830 305, 305 6, 6
103 blosc 1830, 1830 305, 305 6, 6
104 blosc 1830, 1830 305, 305 6, 6
105 blosc 1830 1830 1
106 blosc 1830 1830 1The same information can be retrieved with sf::gdal_utils('mdiminfo', ...).
Reading Zarr data with Rarr
Reading zarrays from a remote Zarr store with Rarr and turning them into useful spatial R objects is not as straightforward as with GDAL-based stars and terra functions. We will demonstrate the current capabilities and limitations of Rarr by using a reference object retrieved through the GDAL Zarr driver which is loaded in the background.
# GDAL-based reference object
zz_gdalstars object with 2 dimensions and 1 attribute
attribute(s), summary of first 1e+05 cells:
Min. 1st Qu. Median Mean 3rd Qu. Max.
b01 0.1299 0.1542 0.1628 0.3541124 0.4304 1.6556
dimension(s):
from to offset delta refsys x/y
x 1 1830 3e+05 60 WGS 84 / UTM zone 32N [x]
y 1 1830 5e+06 -60 WGS 84 / UTM zone 32N [y]We start with an naive attempt to read the entire Zarr store using Rarr::read_zarr_array.
⛔ Rarr, like stars and terra, is not capable of reading multiple arrays from the remote Zarr store simultaneously.
read_zarr_array(zarr_url)Error:
! The requested `.zarray` metadata file (possibly listed in `.zmetadata`) does not exist.✅ But if a single band array is selected we are able to access the desired information.
band_variable = "/measurements/reflectance/r60m/b01"
zarray = read_zarr_array(paste0(zarr_url, band_variable))
str(zarray) int [1:1830, 1:1830] 9548 8117 7158 5906 7007 7523 5999 6570 7165 6666 ...Raster geometry
The function returns a matrix (2D array) lacking any spatial metadata (e.g. Coordinate Reference System) or map coordinates. Let’s plot it together with the reference object for comparison.
image(zarray)
image(zz_gdal$b01)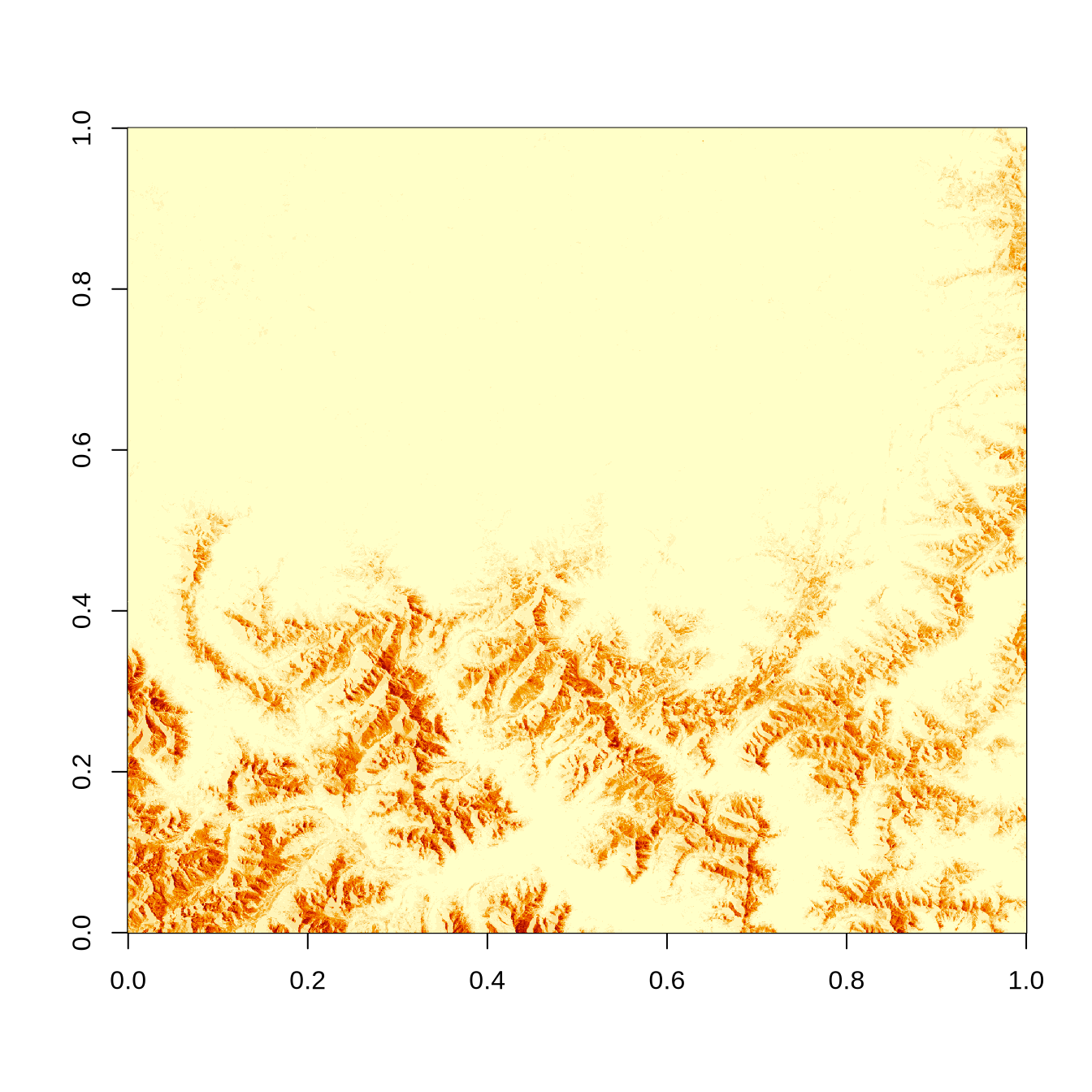
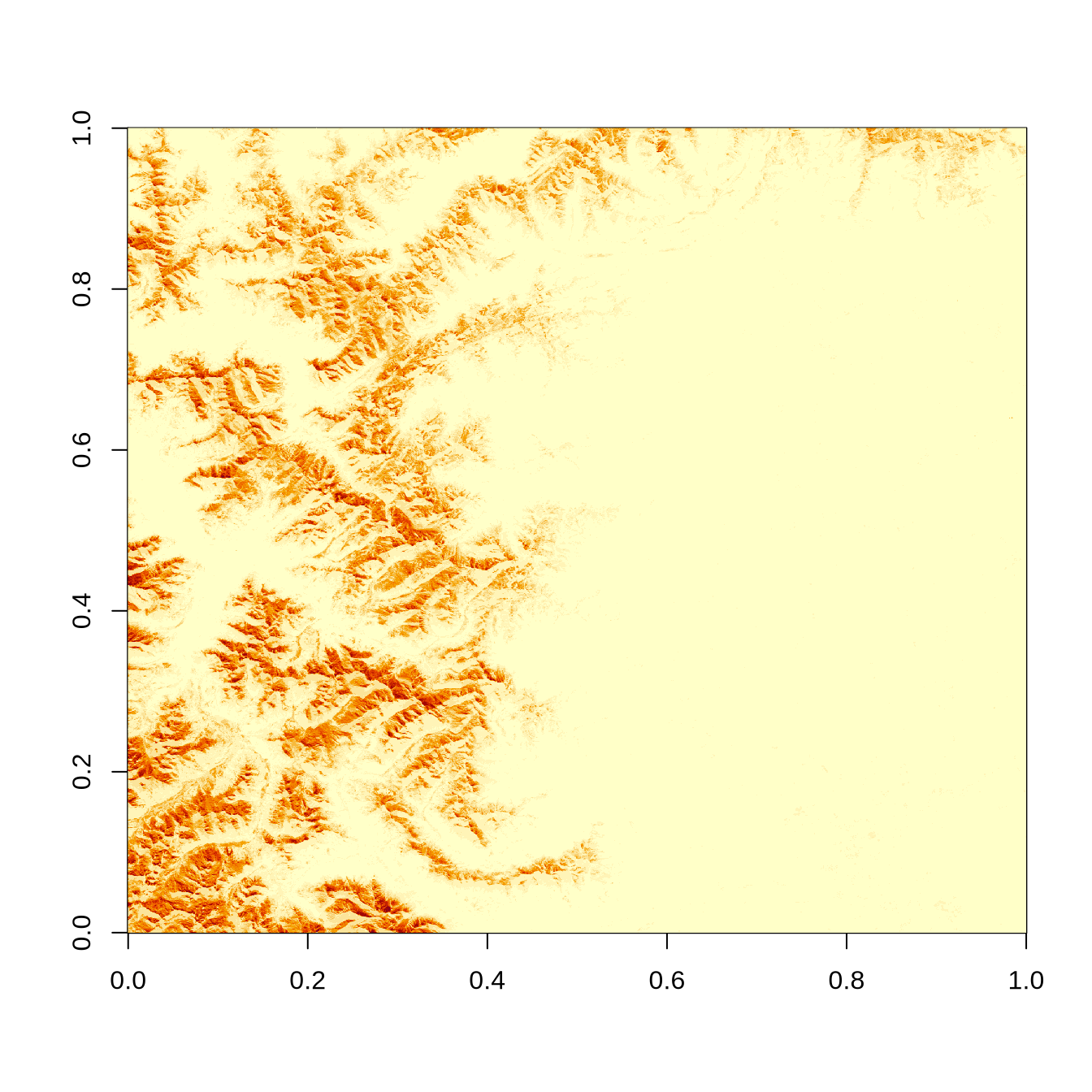
The raw zarray has flipped X- and Y-dimensions! But we can access X and Y coordinates for each pixel separately and handle the issue by transposing them later (using t()).
# retrieve X and Y coordinates
zarray_60m_x = paste0(zarr_url, "/measurements/reflectance/r60m/x") |>
read_zarr_array()
zarray_60m_y = paste0(zarr_url, "/measurements/reflectance/r60m/y") |>
read_zarr_array()
range(zarray_60m_x)[1] 300030 409770range(zarray_60m_y)[1] 4890270 5000010Regardless of their ordering, are the coordinate values consistent with those retrieved via GDAL?
range(st_get_dimension_values(zz_gdal, "x"))[1] 300030 409770range(st_get_dimension_values(zz_gdal, "y"))[1] 4890270 5000010✅ Yes!
Reflectance values
Before building the spatial object we should also check if the actual data values are consistent between both methods.
head(t(zarray)[1,]) # flipped zarray[1] 9548 8117 7158 5906 7007 7523head(zz_gdal$b01[1,]) # reference arrray[1] 0.8548 0.7117 0.6158 0.4906 0.6007 0.6523By comapring the first few values of each array we can observe 2 important differences:
zarraycontains integer values whilezz_gdalhas floating point values scaled with factor 10,000.- after rescaling, min and max for example still differ (0.1299 vs. 0.2299 / 1.6556 vs. 1.7556), indicating an offset of 0.1.
💡 Explanation: The data provider applies scale and offset to store more memory-efficient integer values. A scale factor of 10,000 preserves 4 decimals, while an offset of 0.1 avoids the multiplication of 0. When reading raw data like that we need to apply scale and offset manually to get the original reflectance values.
Scale and offset parameters are important metadata, but currently not accessible via Rarr. At this point we can only find it by digging deep into the metadata retrieved by GDAL:
# GDAL-comaptible VSI URL
vsi_url = paste0("ZARR:/vsicurl/", dplyr::as_label(zarr_url))
# get entire metadata
info = sf::gdal_utils("mdiminfo", source = vsi_url, quiet = T) |> fromJSON()
# access scale and offset for 60 meter reflectance bands
(scale = info$groups$measurements$groups$reflectance$groups$r60m$arrays$b01$scale)[1] 1e-04(offset = info$groups$measurements$groups$reflectance$groups$r60m$arrays$b01$offset)[1] -0.1Constructing a stars object from zarray components
Let’s combine all pieces of the puzzle to form a spatial object:
- data array
- scale and offset
- X and Y coordinates
- CRS
# read CRS from Zarr attributes
zarr_crs = zattrs$stac_discovery$properties$`proj:epsg`
# band name
var = basename(band_variable)
# read and transpose data matrix
# apply scale and offset
# convert to stars
zz = st_as_stars(t(zarray) * scale + offset) |>
st_set_dimensions(1, names = "x", values = zarray_60m_x, is_raster = TRUE) |>
st_set_dimensions(2, names = "y", values = zarray_60m_y, is_raster = TRUE) |>
setNames(var)
# assign CRS
st_crs(zz) = st_crs(zarr_crs)
# print the object
zzstars object with 2 dimensions and 1 attribute
attribute(s), summary of first 1e+05 cells:
Min. 1st Qu. Median Mean 3rd Qu. Max.
b01 0.1299 0.1542 0.1628 0.3541124 0.4304 1.6556
dimension(s):
from to offset delta refsys point x/y
x 1 1830 3e+05 60 WGS 84 / UTM zone 32N FALSE [x]
y 1 1830 5e+06 -60 WGS 84 / UTM zone 32N FALSE [y]We can compare the newly constructed object with reference read via GDAL. They should have identical geometry and values.
identical(zz[[1]] |> str(),
zz_gdal[[1]] |> str()) num [1:1830, 1:1830] 0.855 0.753 0.7 0.756 0.718 ...
num [1:1830, 1:1830] 0.855 0.753 0.7 0.756 0.718 ...[1] TRUELooks good! Let’s visualize both objects.
# plot as maps
plot(zz, axes = TRUE)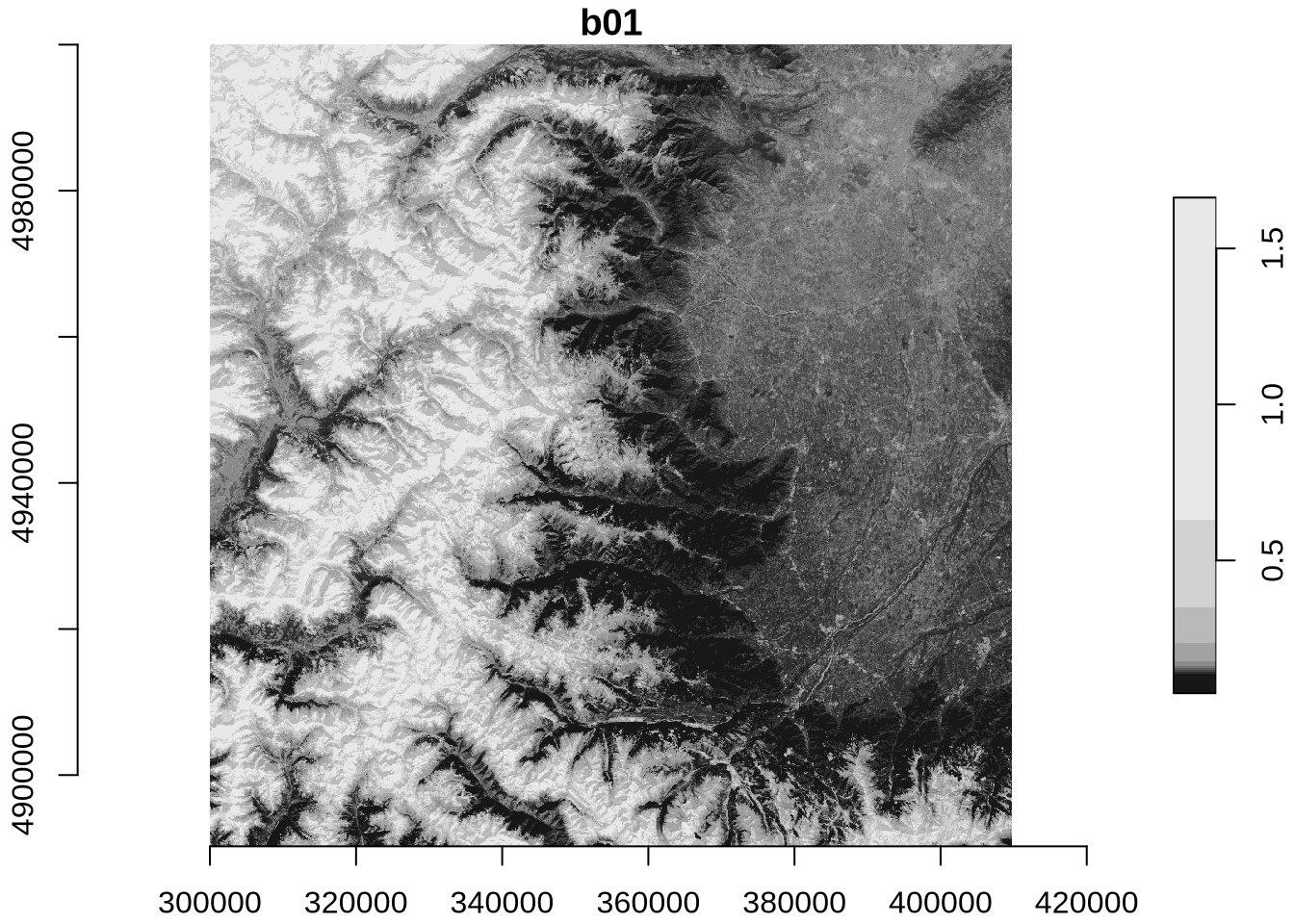
plot(zz_gdal, axes = TRUE)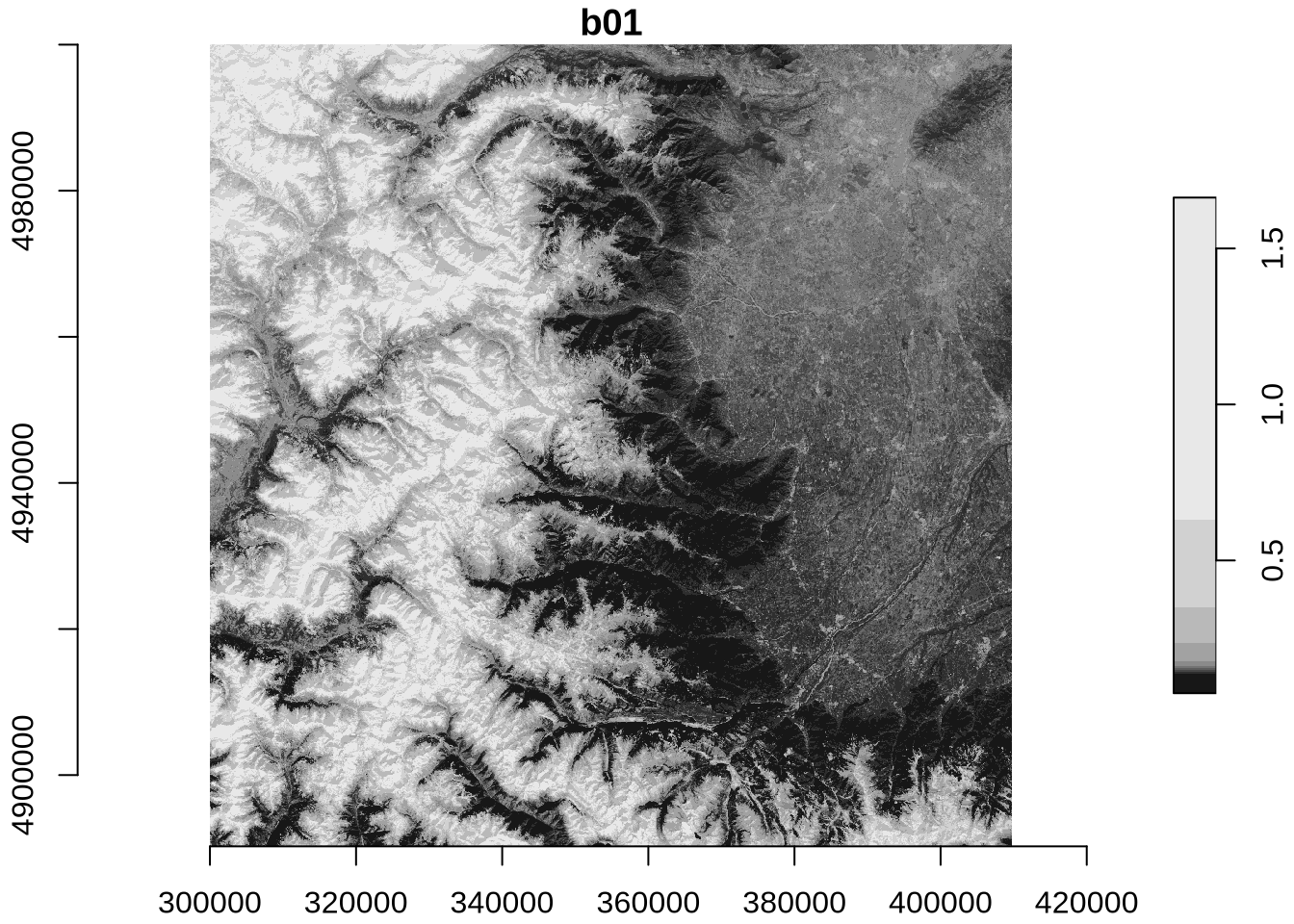
The identical maps show Sentinel-2 band 1 (coastal aerosol band) reflectance at 60m spatial resolution for north-western Italy with the Alps in the West and the city of Torino in the North-East.
Wrapping it all up in a function
We can enclose these steps in a single function with some flexibility for variable names and resolutions. This allows us to retrieve any band at any available resolution from the remote Zarr store. Since scale and offset are not accessible via Rarr, we need to provide them as function arguments.
st_read_zarray = function(path, var, res, scale = 1, offset = 0, ...){
# room for checks and input tests:
# stopifnot valid zarr url / variable / resolution ...
# get metadata including CRS
zattrs = jsonlite::read_json(file.path(path, ".zattrs"))
zarr_crs = zattrs$stac_discovery$properties$`proj:epsg`
# selecte the correct spatial resolution
# zattrs$stac_discovery$properties$`eopf:resolutions`
res_char = switch(as.character(res),
"10" = "r10m",
"20" = "r20m",
"60" = "r60m",
stop("Resolution must be one of 10, 20, or 60.")
)
# get 2D data array and 1D coordinate arrays
# ...: make use of index arguments of read_zarr_array if desired
zarray = file.path(path, "measurements/reflectance", res_char, var) |>
read_zarr_array(...)
zarray_x = file.path(path, "measurements/reflectance", res_char, "x") |>
read_zarr_array(...)
zarray_y = file.path(path, "measurements/reflectance", res_char, "y") |>
read_zarr_array(...)
# transpose matrix before converting to stars, apply scale and offset
# assign x and y coordinates and CRS
z = st_as_stars(t(zarray) * scale + offset) |>
st_set_dimensions(1, names = "x", values = zarray_60m_x, is_raster = TRUE) |>
st_set_dimensions(2, names = "y", values = zarray_60m_y, is_raster = TRUE) |>
setNames(var)
st_crs(z) = st_crs(zarr_crs)
return(z)
}Read and combine multiple zarrays
✅ Now we can read multiple bands, combine them to a single multi-band object, and visualize them in a static map…
system.time({
# scale: 0.0001 = 1 / 10,000
b01 = st_read_zarray(zarr_url, "b01", 60, scale=0.0001, offset = 0.1)
b09 = st_read_zarray(zarr_url, "b09", 60, scale=0.0001, offset = 0.1)
multi_band = c(b01, b09) |> merge(name="band") |> setNames("reflectance")
}) user system elapsed
2.173 0.038 35.519 plot(multi_band, axes = TRUE)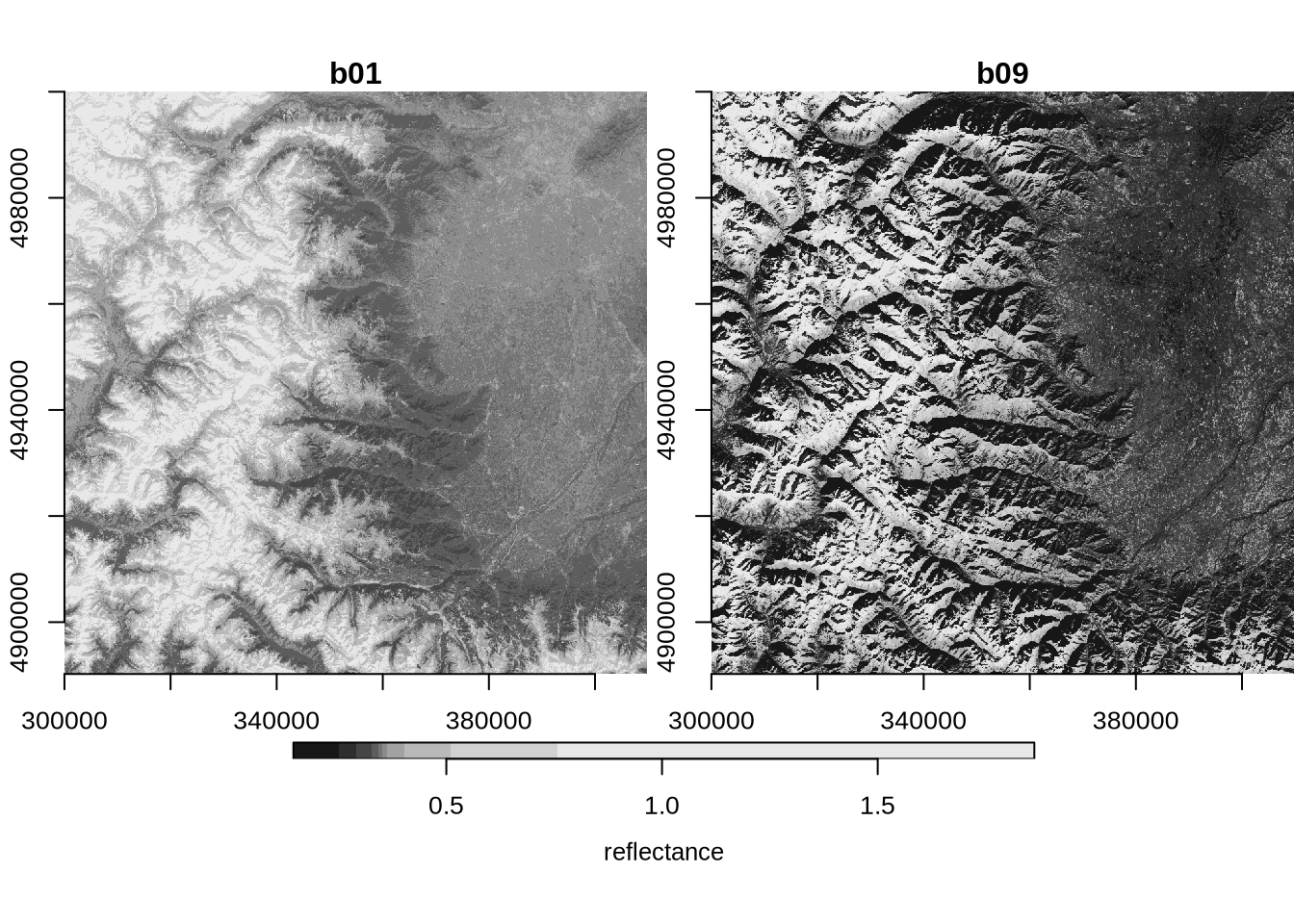
… or by generating an interactive map:
mapview(multi_band)Benchmark
Using Rarr to read remote Zarr stores is just one of several options. R packages stars and terra provide functionality, that uses GDAl’s Zarr driver in the background. To see if there is a benefit to each of the methods, we can compare their execution time and memory allocation with a little benchmark.
# GDAL-compatible VSI URL
vsi_prefix = "ZARR:/vsicurl/"
vsi_url = paste0(vsi_prefix, dplyr::as_label(zarr_url))
# functions for reading a single Sentinel-2 band to memory
f_stars = function(){rs = read_stars(paste(vsi_url, band_variable, sep = ":"))}
f_mdim = function(){sm = read_mdim(vsi_url, band_variable, proxy = FALSE) |>
setNames(basename(band_variable))}
f_rast = function(){tr = terra::rast(vsi_url, band_variable) |> terra::toMemory()}
f_rarr = function(){rr = st_read_zarray(zarr_url, "b01", 60)}bench::mark(f_stars(), f_mdim(), f_rast(), f_rarr(),
check = F, iterations = 5)# A tibble: 4 × 12
expression min median `itr/sec` mem_alloc `gc/sec` n_itr n_gc total_time
<chr> <chr> <chr> <dbl> <chr> <dbl> <int> <dbl> <chr>
1 f_stars() 24s 24.2s 0.0409 25.6MB 0.0163 5 2 2.04m
2 f_mdim() 23.2s 23.3s 0.0427 25.9MB 0.00854 5 1 1.95m
3 f_rast() 23.8s 24s 0.0407 12.2KB 0 5 0 2.05m
4 f_rarr() 13.7s 15.6s 0.0647 147.6MB 0.104 5 8 1.29m
# ℹ 3 more variables: result <list>, time <list>, gc <list>The benchmark does not reveal a clear winner in terms of speed. However, Rarr seems not to be faster than GDAL but allocates more memory compared to stars and terra.
💪 Now it is your turn
- 🔭 Task: Supply more arguments to
st_read_zarrayand pass a subset to read only parts on an array. Visit?Rarr::read_zarr_arrayfor more details. - 💾 Task: Try writing zarrays to disk using
Rarr::write_zarr_array(). - 📖 Read the Rarr documentation.
Conclusion
In this notebook we have demonstrated how to access remote Zarr stores in R using the Rarr package. We have seen that it is possible to read specific data arrays, but read_zarr_array lacks the ability of accessing anything other than the raw array. To combine it with a CRS or coordinates we need to leverage a spatial class like stars and manually set the relevant metadata. Further, the benchmark did not reveal a significant benefit regarding processing time or memory allocation over “traditional” GDAL-based data retrieval.
What’s next?
Further tests should evaluate wether data retrieved via Rarr versus GDAL are identical in terms of spatial extent and offset.
Explore this related notebook which demonstrates how to access the same Zarr data using the GDAL Zarr driver via the stars and terra packages.
